NEST- Nervous system Embryogenesis and Study of paediatric Tumors
Principal investigator: Valérie CASTELLANI
Embryogenesis | developmental biology | pediatric cancers | neuroblastoma | medulloblastoma |cell migration | metastasis | axon guidance | microenvironment | neural crest | enteric nervous system | cortex | spinal cord | avian model | single cell transcriptomic | light sheet microscopy
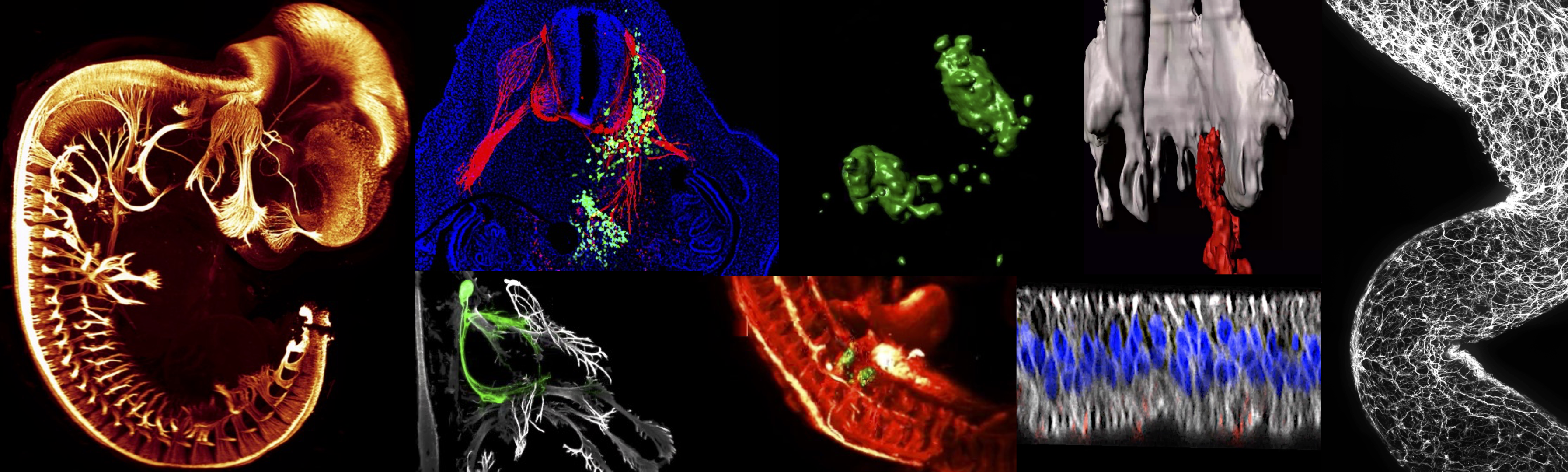
 We study mechanisms underlying embryonic neurodevelopment and malignancies of prenatal origin.
We study mechanisms underlying embryonic neurodevelopment and malignancies of prenatal origin.
Our laboratory studies the cellular and molecular mechanisms that control the formation of the nervous system in the embryo. We are interested in the communication of progenitor and neuronal cells with their environment, with a focus on the cellular processes and molecular signaling that control the colonization of embryonic territories by migrating neural cells and axons. We are currently addressing these questions by studying a remarkable embryonic cell population, the neural crest, at the origin of multiple derivatives, including the sympathetic chain, adrenal medulla, and enteric nervous system. Using experimental manipulations in the avian embryo model and transcriptomics, we study the emergence of enteric neural circuits and brain-gut connectivity to characterize the gene programs that mediate communication between cells and their environment. We are investigating whether these programs are conserved in the human embryo and whether their deregulation might contribute to gut neurodevelopmental disorders.
In parallel, we study pediatric cancers in the light of their embryonic origin, in particular neural crest derived neuroblastoma and cerebellar precursor-derived medulloblastoma. The heterogeneity and plasticity that characterize these malignancies and underlie their aggressiveness are thought to be rooted in the embryonic context of their emergence. Tumorigenic events take place in cells that actively communicate with their environment and are endowed with the proliferative and migratory properties necessary for tissue formation.
While becoming malignant, tumoral cells retain many of the characteristics of the cells of origin. Our goal is to understand how during tumorigenesis and metastasis this dual physiological and tumorigenic potential is manifested by the opportunistic exploitation or hijacking of developmental mechanisms and signaling pathways. This basic research will pave the way for the development of therapies that specifically target tumor-specific behavior and signaling. To address these questions, and taking advantage of our developmental biology models, we have established a specific in vivo paradigm that recapitulates the embryonic context for malignant cells. It consists of transplanting human tumor cells into selected tissues of the avian embryo. Our studies are based on a multi-approach strategy combining experimental embryology, functional studies of genes of interest in avian models, 3D light sheet microscopy to map cells and molecules at the whole embryo level, videomicroscopy, and large-scale transcriptomic analyses.
For the general public
How are our neural circuits built? the newborn neuron develops an extension, the axon, destined to embark on an incredible journey in search of the cells with which it will establish communication. Thus, during embryonic and postnatal development, millions of axons go in search of their partners, some remaining confined to the brain or spinal cord, others colonizing the whole organism to innervate the muscles, the skin, the viscera. Many neuronal cells also migrate to colonize distal territories where they will form a nervous structure. Molecular signals are expressed in embryonic tissues that allow axons and cells to locate themselves in space. They are called topographic or guidance signals. Getting each axon and cell to its destination is a real challenge, and it is these processes that our team is studying. Can axons and cells get lost or misdirected along the way? Alterations of cell and axon migration are responsible for childhood pathologies. Many remain undiscovered because the identification of alterations of these early processes remains difficult. Cells can also become malignant prenatally, at a time when they are undergoing migration and proliferation. They give rise to tumors that can be highly metastatic and have a poor outcome. Our team studies the behavior of malignant cells and how they communicate with embryonic tissues during tumorigenesis and metastasis.
South-ROCK

Our team is a member of the South-ROCK integrated research center of excellence in pediatric oncology of Lyon-Marseille, one of the three laureates of the French National Cancer Institute’s PEDIACRIEX call for projects.
REACT4KIDS

Our team is part of REACT4KIDS (REsearchers in oncology ACTing for kids), a national collaborative network of pediatric oncology research laboratories whose goal is to foster collaborative research for gaining knowledge about the biology of childhood cancers to pave the way for innovative therapies tailored to these cancers.
Start Up

Oncofactory is a start-up founded in 2016 as a spin-off of the Castellani lab, based on the work of Dr. Céline Delloye Bourgeois and Dr. Valérie Castellani at the NeuroMyoGene Institute in Lyon. Oncofactory offers an innovative in vivo platform suited for all cancers consisting in the creation of miniaturized replicas of patient tumors in an embryonic organism. Tumors are analyzed in the whole organism using 3D microscopy. Replicas can be collected back for a range of large scale molecular analysis. Using its unique technological platform Oncofactory evaluates the efficacy, studies the mechanisms of action of therapeutic candidates, helps the design of combi-therapies through a dedicated program combining transcriptomic analysis and its model of patient tumors replicas.
Team Members

- Valérie CASTELLANI — Research Director CNRS, HDR – valerie.castellani@univ-lyon1.fr
- Frédéric MORET — Assistant Professor, UCBL, HDR – frederic.moret@univ-lyon1.fr
- Julien FALK — Researcher, CNRS, HDR – julien.falk@univ-lyon1.fr
- Servane TAUSZIG-DELAMASURE — Research Director, CNRS, HDR – servane.tauszig-delamasure@univ-lyon1.fr
- Muriel BOZON — Research assistant, CNRS – muriel.bozon@univ-lyon1.fr
- Karine THOINET — Research assistant, CNRS – karine.thoinet@univ-lyon1.fr
- Franck BOISMOREAU — Post-doc – franck.boismoreau@univ-lyon1.fr
- Audrey PRUNET — Post-doc – audrey.prunet@univ-lyon1.fr
- Claire RIVET — PhD Student – claire.rivet@univ-lyon1.fr
- Marion MALLET — PhD Student – marion.mallet@univ-lyon1.fr
- Luce ROSEIRO — PhD Student – luce.roseiro@univ-lyon1.fr
- Florian MARTIN — Research Engineer – florian.martin@univ-lyon1.fr
- Maëlys ANDRE — Research Engineer – maelys.andre@univ-lyon1.fr
- Julie PIGNOLET — Research Engineer — julie.pignolet@univ-lyon1.fr
- Baptiste Mida — Research Engineer —
- Myriam BOUDISSA — Research Engineer — myriam.boudissa@univ-lyon1.fr
- Cécile FAURE-CONTER — Oncopediatry Physician IHOPe, CLB – cecile.conter@ihope.fr
- Jérémy GANOFSKY — Research Engineer Bioinformatician – jeremy.ganofsky@univ-lyon1.fr
Alumni
- Céline Delloye-Bourgeois — Group leader, Center of Research of Cancerology of Lyon, France
- Homaira Nawabi — Group leader, Grenoble Institute of Neuroscience, France
- Camille Charoy — Senior microscopist at The Francis Crick Institute, UK
- Florie Reynaud — Research Engineer, Center of Research of Cancerology of Lyon, France
- Elise Arbeille — Assistant professor, University of Méditerranée, Marseille, France
- Anne Briançon-Marjollet — Professor, Grenoble-Alpes University, Grenoble, France
- Aurora Pignata — Assistant professor, University of Toulouse, France
- Leila Boubakar — Post-doc, KU Leuven, Belgium
- Hugo Ducuing — Post-doc, Institut de Recherches Cliniques de Montréal, Canada
- Sarah Dinvaut —Faculty of medecine Rockefeller, Lyon, France
Selected publications
Neuroblastoma plasticity during metastatic progression stems from the dynamics of an early sympathetic transcriptomic trajectory
Villalard B., Boltjes A., Reynaud F., et al.. 🔗 https://doi.org/10.1038/s41467-024-53776-3
Résumé :
Résumé non disponible.
Nature Communications 15, (2024)
GPC3-Unc5 receptor complex structure and role in cell migration
Akkermans O., Delloye-Bourgeois C., Peregrina C., et al.. 🔗 https://doi.org/10.1016/j.cell.2022.09.025
Résumé :
Résumé non disponible.
Cell 185, 3931-3949.e26 (2022)
3D exploration of gene expression in chicken embryos through combined RNA fluorescence in situ hybridization, immunofluorescence, and clearing
André M., Dinvaut S., Castellani V., et al.. 🔗 https://doi.org/10.1186/s12915-024-01922-0
Résumé :
Abstract Background Fine characterization of gene expression patterns is crucial to understand many aspects of embryonic development. The chicken embryo is a well-established and valuable animal model for developmental biology. The period spanning from the third to sixth embryonic days (E3 to E6) is critical for many organ developments. Hybridization chain reaction RNA fluorescent in situ hybridization (HCR RNA-FISH) enables multiplex RNA detection in thick samples including embryos of various animal models. However, its use is limited by tissue opacity. Results We optimized HCR RNA-FISH protocol to efficiently label RNAs in whole mount chicken embryos from E3.5 to E5.5 and adapted it to ethyl cinnamate (ECi) tissue clearing. We show that light sheet imaging of HCR RNA-FISH after ECi clearing allows RNA expression analysis within embryonic tissues with good sensitivity and spatial resolution. Finally, whole mount immunofluorescence can be performed after HCR RNA-FISH enabling as exemplified to assay complex spatial relationships between axons and their environment or to monitor GFP electroporated neurons. Conclusions We could extend the use of HCR RNA-FISH to older chick embryos by optimizing HCR RNA-FISH and combining it with tissue clearing and 3D imaging. The integration of immunostaining makes possible to combine gene expression with classical cell markers, to correlate expressions with morphological differentiation and to depict gene expressions in gain or loss of function contexts. Altogether, this combined procedure further extends the potential of HCR RNA-FISH technique for chicken embryology.
BMC Biology 22, (2024)
Functional precision oncology for follicular lymphoma with patient-derived xenograft in avian embryos
Zala M., Lipinski B., Costechareyre C., et al.. 🔗 https://doi.org/10.1038/s41375-024-02150-9
Résumé :
Résumé non disponible.
Leukemia 38, 430-434 (2024)
Axon guidance during mouse central nervous system regeneration is required for specific brain innervation
Delpech C., Schaeffer J., Vilallongue N., et al.. 🔗 https://doi.org/10.1016/j.devcel.2024.09.005
Résumé :
Résumé non disponible.
Developmental Cell 59, 3213-3228.e8 (2024)
Motor innervation directs the correct development of the mouse sympathetic nervous system
Erickson A., Motta A., Kastriti M., et al.. 🔗 https://doi.org/10.1038/s41467-024-51290-0
Résumé :
AbstractThe sympathetic nervous system controls bodily functions including vascular tone, cardiac rhythm, and the “fight-or-flight response”. Sympathetic chain ganglia develop in parallel with preganglionic motor nerves extending from the neural tube, raising the question of whether axon targeting contributes to sympathetic chain formation. Using nerve-selective genetic ablations and lineage tracing in mouse, we reveal that motor nerve-associated Schwann cell precursors (SCPs) contribute sympathetic neurons and satellite glia after the initial seeding of sympathetic ganglia by neural crest. Motor nerve ablation causes mispositioning of SCP-derived sympathoblasts as well as sympathetic chain hypoplasia and fragmentation. Sympathetic neurons in motor-ablated embryos project precociously and abnormally towards dorsal root ganglia, eventually resulting in fusion of sympathetic and sensory ganglia. Cell interaction analysis identifies semaphorins as potential motor nerve-derived signaling molecules regulating sympathoblast positioning and outgrowth. Overall, central innervation functions both as infrastructure and regulatory niche to ensure the integrity of peripheral ganglia morphogenesis.
Nature Communications 15, (2024)
An in vivo avian model of human melanoma to perform rapid and robust preclinical studies
Jarrosson L., Dalle S., Costechareyre C., et al.. 🔗 https://doi.org/10.15252/emmm.202216629
Résumé :
AbstractMetastatic melanoma patients carrying a BRAFV600 mutation can be treated with a combination of BRAF and MEK inhibitors (BRAFi/MEKi), but innate and acquired resistance invariably occurs. Predicting patient response to targeted therapies is crucial to guide clinical decision. We describe here the development of a highly efficient patient‐derived xenograft model adapted to patient melanoma biopsies, using the avian embryo as a host (AVI‐PDXTM). In this in vivo paradigm, we depict a fast and reproducible tumor engraftment of patient samples within the embryonic skin, preserving key molecular and phenotypic features. We show that sensitivity and resistance to BRAFi/MEKi can be reliably modeled in these AVI‐PDXTM, as well as synergies with other drugs. We further provide proof‐of‐concept that the AVI‐PDXTM models the diversity of responses of melanoma patients to BRAFi/MEKi, within days, hence positioning it as a valuable tool for the design of personalized medicine assays and for the evaluation of novel combination strategies.
EMBO Molecular Medicine 15, (2023)
The Neurod1/4-Ntrk3-Src pathway regulates gonadotrope cell adhesion and motility
Le Ciclé C., Pacini V., Rama N., et al.. 🔗 https://doi.org/10.1038/s41420-023-01615-7
Résumé :
AbstractPituitary gonadotrope cells are essential for the endocrine regulation of reproduction in vertebrates. These cells emerge early during embryogenesis, colonize the pituitary glands and organize in tridimensional networks, which are believed to be crucial to ensure proper regulation of fertility. However, the molecular mechanisms regulating the organization of gonadotrope cell population during embryogenesis remain poorly understood. In this work, we characterized the target genes of NEUROD1 and NEUROD4 transcription factors in the immature gonadotrope αT3-1 cell model by in silico functional genomic analyses. We demonstrated that NEUROD1/4 regulate genes belonging to the focal adhesion pathway. Using CRISPR/Cas9 knock-out approaches, we established a double NEUROD1/4 knock-out αT3-1 cell model and demonstrated that NEUROD1/4 regulate cell adhesion and cell motility. We then characterized, by immuno-fluorescence, focal adhesion number and signaling in the context of NEUROD1/4 insufficiency. We demonstrated that NEUROD1/4 knock-out leads to an increase in the number of focal adhesions associated with signaling abnormalities implicating the c-Src kinase. We further showed that the neurotrophin tyrosine kinase receptor 3 NTRK3, a target of NEUROD1/4, interacts physically with c-Src. Furthermore, using motility rescue experiments and time-lapse video microscopy, we demonstrated that NTRK3 is a major regulator of gonadotrope cell motility. Finally, using a Ntrk3 knock-out mouse model, we showed that NTRK3 regulates gonadotrope cells positioning in the developing pituitary, in vivo. Altogether our study demonstrates that the Neurod1/4-Ntrk3-cSrc pathway is a major actor of gonadotrope cell mobility, and thus provides new insights in the regulation of gonadotrope cell organization within the pituitary gland.
Cell Death Discovery 9, (2023)
A balance of noncanonical Semaphorin signaling from the cerebrospinal fluid regulates apical cell dynamics during corticogenesis
Gerstmann K., Kindbeiter K., Telley L., et al.. 🔗 https://doi.org/10.1126/sciadv.abo4552
Résumé :
During corticogenesis, dynamic regulation of apical adhesion is fundamental to generate correct numbers and cell identities. While radial glial cells (RGCs) maintain basal and apical anchors, basal progenitors and neurons detach and settle at distal positions from the apical border. Whether diffusible signals delivered from the cerebrospinal fluid (CSF) contribute to the regulation of apical adhesion dynamics remains fully unknown. Secreted class 3 Semaphorins (Semas) trigger cell responses via Plexin-Neuropilin (Nrp) membrane receptor complexes. Here, we report that unconventional Sema3-Nrp preformed complexes are delivered by the CSF from sources including the choroid plexus to Plexin-expressing RGCs via their apical endfeet. Through analysis of mutant mouse models and various ex vivo assays mimicking ventricular delivery to RGCs, we found that two different complexes, Sema3B/Nrp2 and Sema3F/Nrp1, exert dual effects on apical endfeet dynamics, nuclei positioning, and RGC progeny. This reveals unexpected balance of CSF-delivered guidance molecules during cortical development.
Science Advances 8, (2022)
Environmental cues from neural crest derivatives act as metastatic triggers in an embryonic neuroblastoma model
Ben Amar D., Thoinet K., Villalard B., et al.. 🔗 https://doi.org/10.1038/s41467-022-30237-3
Résumé :
AbstractEmbryonic malignant transformation is concomitant to organogenesis, often affecting multipotent and migratory progenitors. While lineage relationships between malignant cells and their physiological counterparts are extensively investigated, the contribution of exogenous embryonic signals is not fully known. Neuroblastoma (NB) is a childhood malignancy of the peripheral nervous system arising from the embryonic trunk neural crest (NC) and characterized by heterogeneous and interconvertible tumor cell identities. Here, using experimental models mimicking the embryonic context coupled to proteomic and transcriptomic analyses, we show that signals released by embryonic sympathetic ganglia, including Olfactomedin-1, induce NB cells to shift from a noradrenergic to mesenchymal identity, and to activate a gene program promoting NB metastatic onset and dissemination. From this gene program, we extract a core signature specifically shared by metastatic cancers with NC origin. This reveals non-cell autonomous embryonic contributions regulating the plasticity of NB identities and setting pro-dissemination gene programs common to NC-derived cancers.
Nature Communications 13, (2022)
X-linked partial corpus callosum agenesis with mild intellectual disability: identification of a novel L1CAM pathogenic variant
Bousquet I., Bozon M., Castellani V., et al.. 🔗 https://doi.org/10.1007/s10048-020-00629-y
Résumé :
Résumé non disponible.
neurogenetics 22, 43-51 (2021)
SlitC-PlexinA1 mediates iterative inhibition for orderly passage of spinal commissural axons through the floor plate
Ducuing H., Gardette T., Pignata A., et al.. 🔗 https://doi.org/10.7554/elife.63205
Résumé :
Spinal commissural axon navigation across the midline in the floor plate requires repulsive forces from local Slit repellents. The long-held view is that Slits push growth cones forward and prevent them from turning back once they became sensitized to these cues after midline crossing. We analyzed with fluorescent reporters Slits distribution and FP glia morphology. We observed clusters of Slit-N and Slit-C fragments decorating a complex architecture of glial basal process ramifications. We found that PC2 proprotein convertase activity contributes to this pattern of ligands. Next, we studied Slit-C acting via PlexinA1 receptor shared with another FP repellent, the Semaphorin3B, through generation of a mouse model baring PlexinA1Y1815Fmutation abrogating SlitC but not Sema3B responsiveness, manipulations in the chicken embryo, and ex vivo live imaging. This revealed a guidance mechanism by which SlitC constantly limits growth cone exploration, imposing ordered and forward-directed progression through aligned corridors formed by FP basal ramifications.
eLife 9, (2020)
A Spatiotemporal Sequence of Sensitization to Slits and Semaphorins Orchestrates Commissural Axon Navigation
Pignata A., Ducuing H., Boubakar L., et al.. 🔗 https://doi.org/10.1016/j.celrep.2019.08.098
Résumé :
Résumé non disponible.
Cell Reports 29, 347-362.e5 (2019)
Microenvironment-Driven Shift of Cohesion/Detachment Balance within Tumors Induces a Switch toward Metastasis in Neuroblastoma
Delloye-Bourgeois C., Bertin L., Thoinet K., et al.. 🔗 https://doi.org/10.1016/j.ccell.2017.09.006
Résumé :
Résumé non disponible.
Cancer Cell 32, 427-443.e8 (2017)
Molecular Memory of Morphologies by Septins during Neuron Generation Allows Early Polarity Inheritance
Boubakar L., Falk J., Ducuing H., et al.. 🔗 https://doi.org/10.1016/j.neuron.2017.07.027
Résumé :
Résumé non disponible.
Neuron 95, 834-851.e5 (2017)
Genetic specification of left–right asymmetry in the diaphragm muscles and their motor innervation
Charoy C., Dinvaut S., Chaix Y., et al.. 🔗 https://doi.org/10.7554/elife.18481
Résumé :
The diaphragm muscle is essential for breathing in mammals. Its asymmetric elevation during contraction correlates with morphological features suggestive of inherent left–right (L/R) asymmetry. Whether this asymmetry is due to L versus R differences in the muscle or in the phrenic nerve activity is unknown. Here, we have combined the analysis of genetically modified mouse models with transcriptomic analysis to show that both the diaphragm muscle and phrenic nerves have asymmetries, which can be established independently of each other during early embryogenesis in pathway instructed by Nodal, a morphogen that also conveys asymmetry in other organs. We further found that phrenic motoneurons receive an early L/R genetic imprint, with L versus R differences both in Slit/Robo signaling and MMP2 activity and in the contribution of both pathways to establish phrenic nerve asymmetry. Our study therefore demonstrates L–R imprinting of spinal motoneurons and describes how L/R modulation of axon guidance signaling helps to match neural circuit formation to organ asymmetry.
eLife 6, (2017)
Cerebrospinal fluid-derived Semaphorin3B orients neuroepithelial cell divisions in the apicobasal axis
Arbeille E., Reynaud F., Sanyas I., et al.. 🔗 https://doi.org/10.1038/ncomms7366
Résumé :
Résumé non disponible.
Nature Communications 6, (2015)
PlexinA1 is a new Slit receptor and mediates axon guidance function of Slit C-terminal fragments
Delloye-Bourgeois C., Jacquier A., Charoy C., et al.. 🔗 https://doi.org/10.1038/nn.3893
Résumé :
Résumé non disponible.
Nature Neuroscience 18, 36-45 (2014)
Funding & Support





Past funding






